Scientists Discover a New Signaling Pathway and Design a Novel Drug for Liver Fibrosis
Health & Behavior
Thousands of fluorescent E. coli bacteria make up a biopixel. Photos/Hasty Lab, UC San Diego
In an example of life imitating art, biologists and bioengineers at UC San Diego have created a living neon sign composed of millions of bacterial cells that periodically fluoresce in unison like blinking light bulbs.
Their achievement, detailed in this week’s advance online issue of the journal Nature, involved attaching a fluorescent protein to the biological clocks of the bacteria, synchronizing the clocks of the thousands of bacteria within a colony, then synchronizing thousands of the blinking bacterial colonies to glow on and off in unison.
A little bit of art with a lot more bioengineering, the flashing bacterial signs are not only a visual display of how researchers in the new field of synthetic biology can engineer living cells like machines, but will likely lead to some real-life applications.
Using the same method to create the flashing signs, the researchers engineered a simple bacterial sensor capable of detecting low levels of arsenic. In this biological sensor, decreases in the frequency of the oscillations of the cells’ blinking pattern indicate the presence and amount of the arsenic poison.
Because bacteria are sensitive to many kinds of environmental pollutants and organisms, the scientists believe this approach could be also used to design low cost bacterial biosensors capable of detecting an array of heavy metal pollutants and disease-causing organisms. And because the senor is composed of living organisms, it can respond to changes in the presence or amount of the toxins over time unlike many chemical sensors.
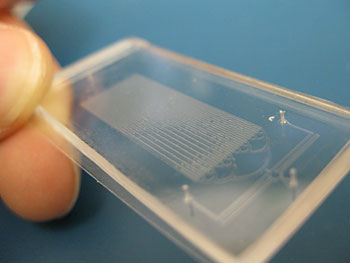

Tiny microfluidic chips allow the researchers to synchronize the bacteria to fluoresce or blink in unison
“These kinds of living sensors are intriguing as they can serve to continuously monitor a given sample over long periods of time, whereas most detection kits are used for a one-time measurement,” said Jeff Hasty, a professor of biology and bioengineering at UC San Diego who headed the research team in the university’s Division of Biological Sciences and BioCircuits Institute. “Because the bacteria respond in different ways to different concentrations by varying the frequency of their blinking pattern, they can provide a continual update on how dangerous a toxin or pathogen is at any one time.”
“This development illustrates how basic, quantitative knowledge of cellular circuitry can be applied to the new discipline of synthetic biology,” said James Anderson, who oversees synthetic biology grants at the National Institutes of Health’s National Institute of General Medical Sciences, which partially funded the research. “By laying the foundation for the development of new devices for detecting harmful substances or pathogens, Dr. Hasty’s new sensor points the way toward translation of synthetic biology research into technology for improving human health.”

The smaller chips contain about 500 blinking bacterial colonies or biopixels
The development of the techniques to make the sensor and the flashing display built on the work of scientists in the Division of Biological Sciences and School of Engineering, which they published in two previous Nature papers over the past four years. In the first paper, the scientists demonstrated how they had developed a way to construct a robust and tunable biological clock to produce flashing, glowing bacteria. In the second paper, published in 2010, the researchers showed how they designed and constructed a network, based on a communication mechanism employed by bacteria, that enabled them to synchronize all of the biological clocks within a bacterial colony so that thousands of bacteria would blink on and off in unison.
“Many bacteria species are known to communicate by a mechanism known as quorum sensing, that is, relaying between them small molecules to trigger and coordinate various behaviors,” said Hasty, explaining how the synchronization works within a bacterial colony. “Other bacteria are known to disrupt this communication mechanism by degrading these relay molecules.”
But the researchers found the same method couldn’t be used to instantaneously synchronize millions of bacteria from thousands of colonies.
“If you have a bunch of cells oscillating, the signal propagation time is too long to instantaneously synchronize 60 million other cells via quorum sensing,” said Hasty. But the scientists discovered that each of the colonies emit gases that, when shared among the thousands of other colonies within a specially designed microfluidic chip, can synchronize all of the millions of bacteria in the chip. “The colonies are synchronized via the gas signal, but the cells are synchronized via quorum sensing. The coupling is synergistic in the sense that the large, yet local, quorum communication is necessary to generate a large enough signal to drive the coupling via gas exchange,” added Hasty.
Graduate students Arthur Prindle, Phillip Samayoa and Ivan Razinkov designed the microfluidic chips, which for the largest ones, contain 50 to 60 million bacterial cells and are about the size of a paper clip or a microscope cover slip. The smaller microfluidic chips, which contain approximately 2.5 million cells, are about a tenth of the size of the larger chips.
Each of the blinking bacterial colonies comprise what the researchers call a “biopixel,” an individual point of light much like the pixels on a computer monitor or television screen. The larger microfluidic chips contain about 13,000 biopixels, while the smaller chips contain about 500 pixels.

The larger chips contain about 13,000 biopixels
Hasty said he believes that within five years, a small hand-held sensor could be developed that would take readings of the oscillations from the bacteria on disposable microfluidic chips to determine the presence and concentrations of various toxic substances and disease-causing organisms in the field.
Other UC San Diego scientists involved in the discovery were Tal Danino and Lev Tsimring.
The UC San Diego Technology Transfer Office has filed a patent application on the Hasty group's invention. Anyone with commercial interest in the research or application should contact Eric Gosink, senior licensing officer, at egosink@ucsd.edu
Keep up with all the latest from UC San Diego. Subscribe to the newsletter today.