Three Generations of UC San Diego Physicists Plumb the Microvasculature of the Mammalian Brain
By:
- Susan Brown
Published Date
By:
- Susan Brown
Share This:
Article Content
Blood vessels within a sensory area of the mammalian brain loop and connect in unexpected ways, a new map created by a team that includes three generations of UC San Diego physicists has revealed.
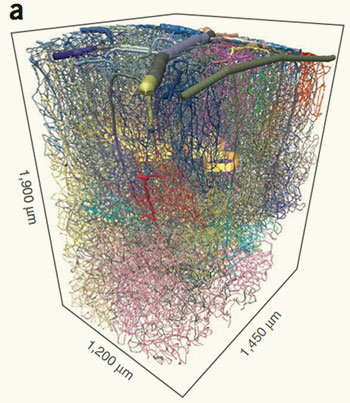
A map of capillaries in the mouse cerebral cortex. Golden bands encircle columns of cells that respond to wiggles of individual whiskers. Other colors mark groups of vessels that are closely connected.
The study, published June 9 in the early online edition of Nature Neuroscience, describes vascular architecture within a well-known region of the cerebral cortex and explores what that structure means for functional imaging of the brain and the onset of a kind of dementia.
David Kleinfeld (Ph.D. ’84), professor of physics and neurobiology at UC San Diego, and colleagues mapped blood vessels in an area of the mouse brain that receives sensory signals from the whiskers.
The brain area is part of the cerebral cortex, which is fed by small arteries that plunge from the surface of the brain and is drained by small veins that return from the depths to the surface. The network of capillaries in between was uncharted though these tiny vessels deliver crucial oxygen and nutrients to energy-hungry brain cells and carry away wastes.
The team traced this fine network by filling the vessels with a fluorescent gel. Then, using an automated system, developed by co-author Philbert Tsai (B.S. ’94, Ph.D. ’04), a project scientist in physics, reconstructed the three-dimensional network of tiny vessels. The system, which removes thin layers of tissue with a laser while capturing a series of images, will be instrumental to the rapid mapping planned by the recently announced BRAIN initiative.
This project focused on a region of the cerebral cortex in which the nerve cells are so well known they can be traced to individual whiskers. These neurons cluster in “barrels,” one per whisker, a pattern of organization seen in other sensory areas as well.
The scientists expected each whisker barrel to match up with its own discrete blood supply, but that was not the case.
“This was a surprise, because the blood vessels develop in tandem with neural tissue,” Kleinfeld said. Instead, microvessels beneath the surface loop and connect in patterns that don't obviously correspond to the barrels.
To search for hidden patterns, they turned to a branch of mathematics called graph theory, which describes systems as connecting paths. Using this approach, led by Harry Suhl, professor emeritus of physics and a founding faculty member who joined the campus in 1961, they determined that the mesh indeed forms a continous network they call the “angiome.”
The vascular maps traced in this study raise a question of what we're actually seeing in a widely used kind of brain imaging called functional MRI, which in one form measures brain activity by recording changes in oxygen levels in the blood. The idea is that activity will locally deplete oxygen. But without a discrete blood supply, they wondered how precisely that optical signal would match the sites of neural activity.
To find out, they wiggled whiskers on individual mice and found that optical signals associated with depleted oxygen indeed centered on the barrels, where electrical recordings confirmed neural activity.
The researchers also went a step further to calculate patterns of blood flow based on the diameters and connections of the vessels and asked how this would change if a feeder arteriole were blocked. The map allowed them to identify "perfusion domains," which predict the volumes of lesions that result when a clot occludes a vessel. Critically, they were able to build a physical model of how these lesions form, as may occur in cases of human vascular dementia.
Additional co-authors include Pablo Blinder, John Kaufhold and Per Knutsen. This work was funded by the National Institutes of Health, including a Director's Pioneer Award to Kleinfeld.
Share This:
You May Also Like
Stay in the Know
Keep up with all the latest from UC San Diego. Subscribe to the newsletter today.


