‘SUMO’ Research at UC San Diego Wrestles with New Way to Read Cellular Function
By:
- Cynthia Dillon
- Mario Aguilera
Media Contact:
- Cynthia Dillon - cdillon@ucsd.edu
- Mario Aguilera - maguilera@ucsd.edu
Published Date
By:
- Cynthia Dillon
- Mario Aguilera
Share This:
Article Content
University of California San Diego researchers in the Department of Chemistry and Biochemistry and in the Section of Cell and Developmental Biology have “got mail”—of the cellular sort. They know that cells have elaborate “addressing” functions that “send” proteins to the correct compartment, but they are now learning how cells “’write” the addresses and then “read” them. This is important because cellular function depends on each molecule in the cell being where it is supposed to be. But, often, protein molecules are not made in the compartment where they eventually need to function. In a paper recently published in Nature Communications, Professor Elizabeth Komives, chemistry and biochemisty, and Associate Professor Eric Bennett, cell and developmental biology, outline their research.
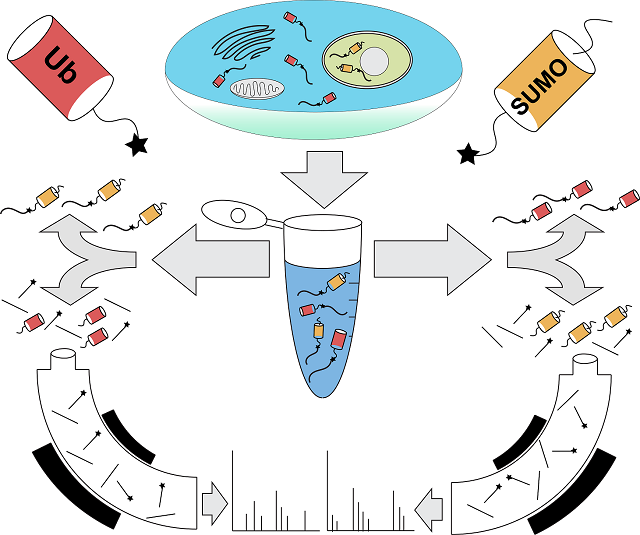
According to the scientists, ubiquitin and SUMO (which stands for Small Ubiquitin-like MOdifier) are protein labels that function in the cell like addresses, telling the cell where to send the labeled protein. Bennett was part of the team at Harvard that figured out how to read the ubiquitin address, and that is why Komives approached him to collaborate with her when she had the idea of a way to read the SUMO address. Together, along with their team of researchers, who included Ryan Lumpkin, doctoral graduate student; Marilyn Leonard, staff research associate and Jesse Meyer, doctoral graduate student, the collaborative colleagues developed a new methodology by which to read both the ubiquitin and SUMO addresses simultaneously.
Previously, researchers generally lacked the tools to be able to identify the proteins that are modified by SUMO from a native cell or tissue source. The Bennet-Komives research, however, opens up a floodgate of new studies to figure out what happens to proteins that are targeted by SUMO. Examples of target proteins include key factors that regulate transcription, protein localization to the nucleus and the response to damaged DNA.
“We have developed a new method to detect, simultaneously, two disease-relevant modifications on proteins from any cell or tissue source,” explained Bennett. “This study represents the first description of the natural targets of one of these modifications, without requiring protein or cell engineering.”
These findings allow other scientists to examine which proteins are “sumoylated” and how that changes during development, exposure to environmental stresses or during disease development. The new methodology involves cutting proteins into smaller portions using a specialized enzyme developed in Komives’ lab. The subset of these small portions that contain a unique signature from SUMO can be captured and subsequently identified by mass spectrometry—an analytical technique that measures mass-to-charge ratios within ionized chemicals.
“This tool can be further refined to gain a deeper appreciation for the types of pathways regulated by SUMO. Known tumor suppressors and oncogenes are modified by SUMO. We are now poised to investigate how protein sumoylation on key disease targets is altered during a time course of disease progression,” said Bennett.
The research, “Site-specific identification and quantitation of endogenous SUMO modifications under native conditions,” was supported by NSF (grant MCB 1244506), NIH (grants T32 EB009380, T32 GM 008326 and T32 GM 008326), New Scholar awards from the Sidney Kimmel Foundation for Cancer Research and the Ellison Medical Foundation, a Hellman Fellowship and a NIH New Innovator Award (grant DP2 GM119132).
The Divisions of Physical Sciences and Biological Sciences at UC San Diego will get an added boost in fall 2018 when the Tata Hall for the Sciences opens with new collaborative research spaces.
Share This:
You May Also Like
Stay in the Know
Keep up with all the latest from UC San Diego. Subscribe to the newsletter today.