
Virus-Like Probes Could Help Make Rapid COVID-19 Testing More Accurate, Reliable
Published Date
By:
- Liezel Labios
Share This:
Article Content
Nanoengineers at the University of California San Diego have developed new and improved probes, known as positive controls, that could make it easier to validate rapid, point-of-care diagnostic tests for COVID-19 across the globe.
The positive controls, made from virus-like particles, are stable and easy to manufacture. Researchers say the controls have the potential to improve the accuracy of new COVID-19 tests that are simpler, faster and cheaper, making it possible to expand testing outside the lab.
“Our goal is to make an impact not necessarily in the hospital, where you have state-of-the-art facilities, but in low-resource, underserved areas that may not have the sophisticated infrastructure or trained personnel,” said Nicole Steinmetz, a professor of nanoengineering at the UC San Diego Jacobs School of Engineering.
Positive controls are a staple in the lab—they are used to verify that a test or experiment indeed works. The positive controls that are primarily used to validate today’s COVID-19 tests are naked synthetic RNAs, plasmids or RNA samples from infected patients. But the issue is RNA and plasmids are not stable like viral particles. They can degrade easily and require refrigeration, making them inconvenient and costly to ship around the world or store for long periods of time.
In a paper published Nov. 25 in ACS Nano, UC San Diego researchers led by Steinmetz report that by packaging segments of RNA from the SARS-CoV-2 virus into virus-like particles, they can create positive controls for COVID-19 tests that are stable—they can be stored for a week at temperatures up to 40 C (104 F), and retain 70% of their activity even after one month of storage—and can pass detection as the novel coronavirus without being infectious.
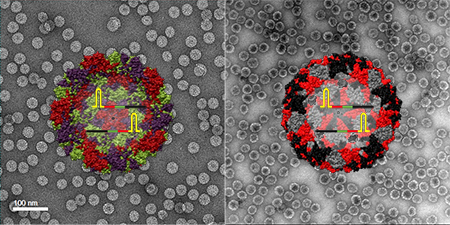
Illustration and TEM image of SARS-CoV-2 positive control made from plant virus-based nanoparticles (left) and bacteriophage nanoparticles (right). Image courtesy of Soo Khim Chan/ACS Nano
The team developed two different controls: one made from plant virus nanoparticles, the other from bacteriophage nanoparticles. Using them is simple. The controls are run and analyzed right alongside a patient sample, providing a reliable benchmark for what a positive test result should look like.
To make the plant virus-based controls, the researchers use the cowpea chlorotic mottle virus, which infects black-eyed pea plants. They essentially open the virus, remove its RNA contents, replace them with a synthesized RNA template containing specific sequences from the SARS-CoV-2 virus, then close everything back up.
The process to make the bacteriophage-based controls starts with plasmids, which are rings of DNA. Inserted into these plasmids are the gene sequences of interest from the SARS-CoV-2 virus, as well as genes coding for surface proteins of the bacteriophage Qbeta. These plasmids are then taken up by bacteria. This process reprograms the bacteria to produce virus-like particles with SARS-CoV-2 RNA sequences on the inside and Qbeta bacteriophage proteins on the outside.
Both controls were validated with clinical samples. A big advantage, the researchers point out, is that unlike the positive controls used today, these can be used in all steps of a COVID-19 test.
“We can use these as full process controls—we can run the analysis in parallel with the patient sample starting all the way from RNA extraction,” said first author Soo Khim Chan, a postdoctoral researcher in Steinmetz’s lab. “Other controls are usually added at a later step. So if something went wrong in the first steps, you won’t be able to know.”
So far, the researchers have adapted their controls for use in the CDC-authorized RT-PCR test. While this is currently the gold standard for COVID-19 testing, it is expensive, complex, and can take days to return results due to the logistics of sending samples off to a lab with PCR capability.
Steinmetz, Chan and colleagues are now working on adapting the controls for use in less complex diagnostic tests like the RT-LAMP test that can be done on the spot, out of the lab and provide results right away.
“It’s a relatively simple nanotechnology approach to make low-tech assays more accurate,” Steinmetz said. “This could help break down some of the barriers to mass testing of underserved populations in the U.S. and across the world.”
Paper title: “Biomimetic Virus-like Particles as Severe Acute Respiratory Syndrome Coronavirus 2 Diagnostic Tools.”
This work was funded in part by grants from the National Science Foundation: RAPID CBET-2032196 and RAPID CMMI-2027668, as well as the University of California: UCOP-R00RG2471 and a Galvanizing Engineering in Medicine (GEM) Award.
Share This:
You May Also Like
Stay in the Know
Keep up with all the latest from UC San Diego. Subscribe to the newsletter today.


