What Molecules You Leave on Your Phone Reveal About Your Lifestyle
Your diet, medications and beauty products leave molecular traces on the objects you touch, providing an unbiased, data-driven profiling method for crime scene investigations and other potential applications
Published Date
By:
- Heather Buschman
Share This:
Article Content
We leave behind trace chemicals, molecules and microbes on every object we touch. By sampling the molecules on cell phones, researchers at University of California San Diego School of Medicine and Skaggs School of Pharmacy and Pharmaceutical Sciences were able to construct lifestyle sketches for each phone’s owner, including diet, preferred hygiene products, health status and locations visited. This proof-of-concept study, published November 14 by Proceedings of the National Academy of Sciences, could have a number of applications, including criminal profiling, airport screening, medication adherence monitoring, clinical trial participant stratification and environmental exposure studies.
“You can imagine a scenario where a crime scene investigator comes across a personal object — like a phone, pen or key — without fingerprints or DNA, or with prints or DNA not found in the database. They would have nothing to go on to determine who that belongs to,” said senior author Pieter Dorrestein, PhD, professor in UC San Diego School of Medicine and Skaggs School of Pharmacy and Pharmaceutical Sciences. “So we thought — what if we take advantage of left-behind skin chemistry to tell us what kind of lifestyle this person has?”
In a 2015 study, Dorrestein’s team constructed 3D models to illustrate the molecules and microbes found at hundreds of locations on the bodies of two healthy adult volunteers. Despite a three-day moratorium on personal hygiene products before the samples were collected, the researchers were surprised to find that the most abundant molecular features in the skin swabs still came from hygiene and beauty products, such as sunscreen.
“All of these chemical traces on our bodies can transfer to objects,” Dorrestein said. “So we realized we could probably come up with a profile of a person’s lifestyle based on chemistries we can detect on objects they frequently use.”
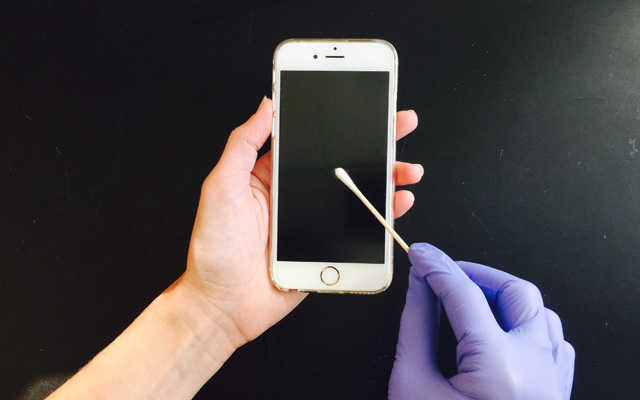
Thirty-nine healthy adult volunteers participated in Dorrestein’s latest study. The team swabbed four spots on each person’s cell phone — an object we tend to spend a lot of time touching — and eight spots on each person’s right hand, for a total of nearly 500 samples. Then they used a technique called mass spectrometry to detect molecules from the samples. They identified as many molecules as possible by comparing them to reference structures in the GNPS database, a crowdsourced mass spectrometry knowledge repository and annotation website developed by Dorrestein and co-author Nuno Bandeira, PhD, associate professor at the Jacobs School of Engineering and Skaggs School of Pharmacy and Pharmaceutical Sciences at UC San Diego.
With this information, the researchers developed a personalized lifestyle “read-out” from each phone. Some of the medications they detected on phones included anti-inflammatory and anti-fungal skin creams, hair loss treatments, anti-depressants and eye drops. Food molecules included citrus, caffeine, herbs and spices. Sunscreen ingredients and DEET mosquito repellant were detected on phones even months after they had last been used by the phone owners, suggesting these objects can provide long-term composite lifestyle sketches.
“By analyzing the molecules they’ve left behind on their phones, we could tell if a person is likely female, uses high-end cosmetics, dyes her hair, drinks coffee, prefers beer over wine, likes spicy food, is being treated for depression, wears sunscreen and bug spray — and therefore likely spends a lot of time outdoors — all kinds of things,” said first author Amina Bouslimani, PhD, an assistant project scientist in Dorrestein’s lab. “This is the kind of information that could help an investigator narrow down the search for an object’s owner.”
There are limitations, Dorrestein said. First of all, these molecular read-outs provide a general profile of person’s lifestyle, but they are not meant to be a one-to-one match, like a fingerprint. To develop more precise profiles and for this method to be more useful, he said more molecules are needed in the reference database, particularly for the most common foods people eat, clothing materials, carpets, wall paints and anything else people come into contact with. He’d like to see a trace molecule database on the scale of the fingerprint database, but it’s a large-scale effort that no single lab will be able to do alone.
Moving forward, Dorrestein and Bouslimani have already begun extending their study with an additional 80 people and samples from other personal objects, such as wallets and keys. They also hope to soon begin gathering another layer of information from each sample — identities of the many bacteria and other microbes that cover our skin and objects. In a 2010 study, their collaborator and co-author, Rob Knight, PhD, professor in the UC San Diego School of Medicine and Jacobs School of Engineering and director of the Center for Microbiome Innovation at UC San Diego, contributed to a study in which his team found they could usually match a computer keyboard to its owner just based on the unique populations of microbes the person left on it. At that time, they could make the match with a fair amount of accuracy, though not yet precisely enough for use in an investigation.
Beyond forensics, Dorrestein and Bouslimani imagine trace molecular read-outs could also be used in medical and environmental studies. For example, perhaps one day physicians could assess how well a patient is sticking with a medication regimen by monitoring metabolites on his or her skin. Similarly, patients participating in a clinical trial could be divided into subgroups based on how they metabolize the medication under investigation, as revealed by skin metabolites — then the medication could be given only to those patients who can metabolize it appropriately. Skin molecule read-outs might also provide useful information about a person’s exposure to environmental pollutants and chemical hazards, such as in a high-risk workplace or a community living near a potential pollution source.
Study co-authors also include: Alexey V. Melnik, Zhenjiang Zech Xu, Amnon Amir, Ricardo R. da Silva, Mingxun Wang, UC San Diego; and Theodore Alexandrov, UC San Diego, European Molecular Biology Laboratory and SCiLS GmbH.
This research was funded, in part, by the National Institute of Justice (2015-DN-BX-K047), National Institutes of Health (GMS10RR029121), European Union’s Horizon 2020 Programme (634402) and São Paulo Research Foundation (FAPESP-2015/03348-3).
Share This:
You May Also Like
Stay in the Know
Keep up with all the latest from UC San Diego. Subscribe to the newsletter today.