
By:
- Heather Buschman
Published Date
By:
- Heather Buschman
Share This:
How to Spot Patients Most Likely to Die from Blood Infections
David Gonzalez’s “a-ha” moment came when a physician-colleague, George Sakoulas, MD, shared with him one of the biggest problems faced in clinical practice: How long it takes to diagnose a patient.
“The faster we know what’s going to happen to our patients, the better we can treat them,” said Sakoulas, an infectious disease specialist and associate adjunct professor of pediatrics at University of California San Diego School of Medicine.
Gonzalez is a biochemist who specializes in proteomics. As genomics is the study of all the genes in a cell or organism, proteomics is the study of all of the proteins. He uses leading-edge tools to identify the proteins in a mixed sample based on their molecular weights—a technique called mass spectrometry.
UC San Diego researchers uncovered biomarkers associated with a patient’s risk of dying from Staphylococcus aureus bacteremia—a leap forward toward a predictive tool that would allow clinicians to more quickly determine which patients require a more complex treatment regimen.
So Gonzalez thought: What if a proteomic “readout” from a person’s blood could help identify who needs the most help early on, so they can be treated quickly and appropriately?
Now, just two years since receiving their first patient blood samples for study, Gonzalez and colleagues have identified a collective signature of proteins and metabolites associated with death due to Staphylococcus aureus bacteremia—a bacterial infection in the blood that kills 20-30% of patients who contract it. In the lab, scientists say these molecular indicators, or biomarkers, can predict who is at highest risk of dying from the infection with exceptional accuracy.
In the study, published Sept. 3, 2020 in Cell, the team describes one of the most comprehensive molecular assessments of blood serum from any human infection response to date. They also validated their findings in mouse models of S. aureus bacteremia.
“This finding is a leap forward toward a point-of-care predictive tool for bacteremia risk,” said Gonzalez, PhD, senior author and assistant professor at UC San Diego School of Medicine and Skaggs School of Pharmacy and Pharmaceutical Sciences. “It also opens up lots of new basic biological questions about how our immune systems respond to infections.” Gonzalez led the study with first author Jacob Wozniak, PhD, a graduate student in his lab at the time.
Gonzalez and team used mass spectrometry to analyze more than 10,000 proteins and metabolites in more than 200 serum samples collected from the blood of patients with Staphylococcus aureus bacteremia. Serum is notoriously difficult to study, he said, because it is heavily laden with a handful of highly abundant serum proteins.
“So, at first, the depth of our proteomic data was a total let down,” Gonzalez said. “We didn’t learn as much as we had hoped about the serum proteins.”
But that initial hurdle only inspired the team to look deeper, at post-translational modifications—the chemical additions added to proteins after they are constructed. According to Gonzalez, post-translational modifications are mostly uncharted territory. Many research efforts are geared toward genomics, but the gene that encodes a protein doesn’t reveal much about how it might be modified later.
“If I wanted to learn all about you, I’d just talk to you directly, not your second cousin,” Gonzalez said. “Same thing here—we can gain new and important information by directly ‘asking’ the proteins, rather than their genes, and mass spectrometry is currently the best way to do that in an unbiased manner.”
With this approach, the team identified a specific pattern of proteins with and without post-translational modifications that differed in the serum of patients who ultimately died of S. aureus bacteremia compared to those who did not. The biomarkers most highly associated with death included lower levels of glycosylated (sugar-coated) fetuin A, unmodified fetuin B and thyroxine, a master regulator of metabolism, as well as higher levels of serum protein carbamylation, another post-translational modification.
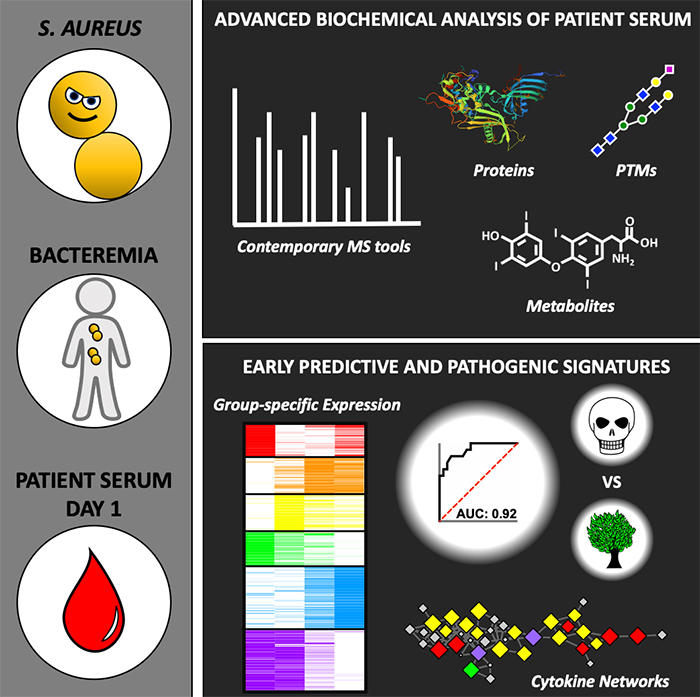
Using advanced biochemical analysis of patient serum, UC San Diego researchers found signatures that help predict which patients are at highest risk of dying from Staphylococcus aureus bacteremia, a blood infection.
Several of these new biomarkers are already known to be associated with disease—high fetuin levels are associated with obesity and diabetes, carbamylation has been linked with kidney disease—but few have been previously linked to bacterial infections.
While the analyses revealed serum differences between low- and high-risk patients, it wasn’t clear whether these molecules actually contribute to the disease, or are simply bystanders. So Gonzalez and team used a mouse model of S. aureus bacteremia to explore cause and effect. They found that mice with higher thyroxine levels had a four-times greater survival rate at 48 hours after infection than control mice. These results indicated that at least one of the identified biomarkers plays a direct role in disease outcome.
In the past, other research groups have developed alternative methods for predicting a patient’s risk of death due to bacteremia. At best, Gonzalez says their accuracy was fair to good. With his team’s new, proteomics-based prediction method, they could predict who is most likely to die of S. aureus bacteremia with excellent predictability. To put it quantitatively, the area under the curve (AUC) was 0.95; 1.0 is perfect and anything above 0.90 is considered excellent in this standard measure of the ability of a test to correctly classify those with and without the disease.
“We tend to treat all bacteremia patients with the same cheap antibiotics, yet we know they only work for 80% of these patients,” said Sakoulas, a co-author of the study. “We need to know from the beginning who falls into that 20% that will require a more complex treatment regimen, so we don’t waste time with trial-and-error.”
Now the team is working to translate their mass spectrometry observations in the laboratory into a rapid clinical test that uses antibody probes to detect S. aureus bacteremia-associated proteins. They are also expanding the approach to look at proteomic and metabolomic markers indicative of high-risk patients with other types of infections, including COVID-19. In addition, researchers are following up on the proteins and modifications that were revealed in the study, exploring their origins, roles in the immune response and potential as therapeutic targets.
Share This:
You May Also Like
Stay in the Know
Keep up with all the latest from UC San Diego. Subscribe to the newsletter today.


