
SDSC’s New ‘SeedMe’ Resource Invites Researchers to Rapidly Share Results
NSF-Funded Project Stresses Ease-of Use and Automation
Published Date
By:
- Jan Zverina
Share This:
Article Content
In late 2012, researchers at the San Diego Supercomputer Center (SDSC) at the University of California, San Diego, were awarded a three-year, $810,000 grant from the National Science Foundation to develop a resource that lets scientists seamlessly share and stream data-intensive visualizations on a variety of platforms, including mobile devices.
Fast forward to the present, where the team of SDSC researchers has developed a working implementation of SeedMe, short for ‘Swiftly Encode, Explore, Disseminate My Experiments.’ The team developed a web-based architecture that not only supports videos and other visualizations, but content such as plots, files, and tickers – all of which are essential to scientific research. The goal is to enable rapid sharing of content directly from applications running on HPC- or cloud-based resources.

“SeedMe provides an essential yet missing component in current high-performance computing,” said Amit Chourasia, a senior visualization scientist at SDSC and principal investigator for the project. “We are now inviting researchers from industry and academia, and across numerous domains, to familiarize themselves with SeedMe’s easy-to-use functionality for sharing science results with their peers or the public at large,”
Easy-to-use programmatic and command line tools are available with documentation for end-users to upload and share their content at SeedMe.org. For a swift introduction a quick start guide, is also available. Moreover, users can easily upload and edit the content using the Web Browser.
“The current HPC infrastructure relies heavily on controlled access,” said Michael Norman, SDSC’s director and co-principal investigator for the project. “SeedMe provides secure input and public output complementing HPC resources without compromising their security model. We believe this project will have a transformative impact throughout numerous domains, as research has become increasingly collaborative, requiring better tools and resources that aid the sharing of information.”

Key features of the SeedMe platform
Unique Solution for the Research Community
Current methods for sharing and assessing results can be labor-intensive and unsupported by useful tools and procedures. Typically, a research team must create their own scripts and ad hoc procedures just to move results from system to system or user to user. These efforts usually rely on email, ftp, or secure copy (scp) despite the ubiquity of more flexible web-based technologies, and do not fully take advantage of the interactive capabilities of today’s mobile devices.
SeedMe is unique from other web-based interfaces in that it is completely focused on efficiently sharing scientific content. While SeedMe could be described in general terms as a DropBox for science, the computational research community requires several additional capabilities and integration tools.
One such requirement is easy instrumentation of simulation codes, as well as support for instrumenting analysis routines from a variety of software tools. That instrumentation may provide periodic tracking and monitoring by posting progress messages, job status, early data snippets, scripts, parameter files, and other reusable components.
Another key requirement is an ability to describe content such as plots and meta data about the research content. This can be accomplished easily with SeedMe.
“SeedMe aims to fill the automation and accessibility gap in computational research,” said Chourasia. “We believe that no single web service or tool suite addresses the breadth of scientific computing challenges as effectively and easily as the SeedMe infrastructure.”
Visualization Speed-ups
Visualizations and simulations are an ever growing part of today’s scientific research. These state-of-the-art simulations are now creating visualization images at runtime using in-situ processing. Images or sequence of images can now be shared with others using SeedMe with simple asynchronous integration, providing an easy monitoring and assessment mechanism for collaborators.
SeedMe also capitalizes on recent strides in video standardization, by encoding videos at very high quality with several resolutions and at desired frame rate that is necessary for research content and also making these available for download to researchers.
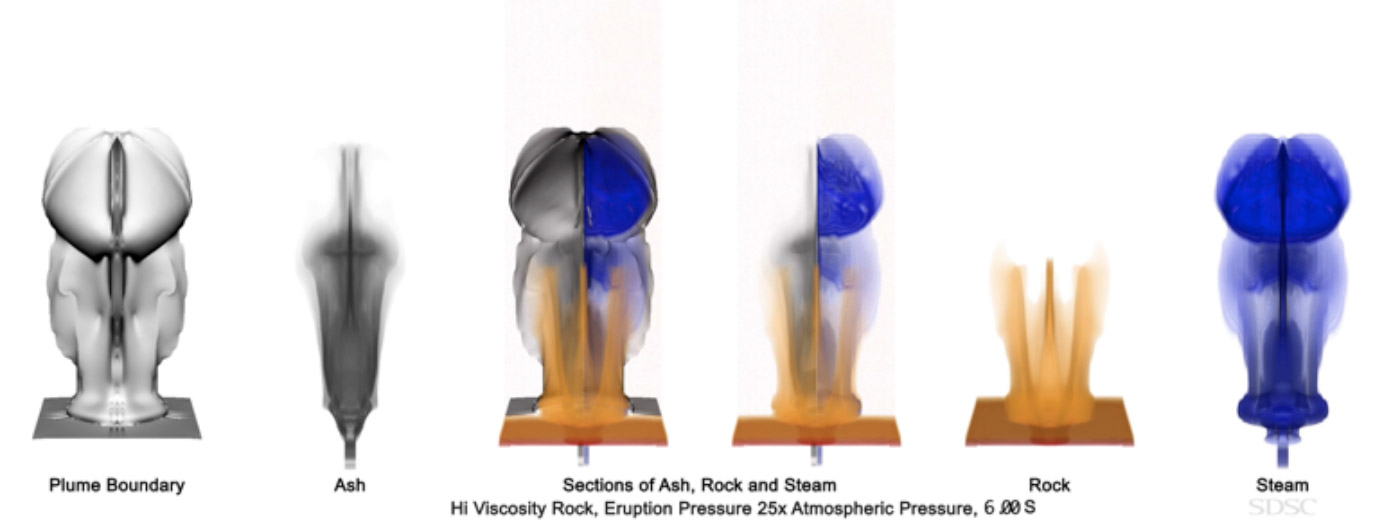
An image from a sequence of visualizations created from data of volcano eruption simulation, showing distribution of various components. Source: Amit Chourasia and Darcy Ogden UC San Diego.
Videos and images are available at SeedMe.
“Our goal has been to convert a manual, serial, error-prone process into an automatic, easy-to-use streamlined, parallel and web accessible cyberinfrastructure, and at the same time create videos that are not only encoded at very high quality to mitigate compression artifacts for scientific content, but also make these easily downloadable for researchers” said Chourasia. “This automation means that students, scientists, and the research community at large does not always have to go through the time and pain to properly encode visualization movies that can be played back on any computer or mobile device.”
Early Adopters
Early users are welcoming SeedMe, which Chourasia envisions as the “go-to place” for sharing scientific content and results. “It is so convenient to share research results with collaborators, compared with sending results through emails,” said Hao Xu, a post-doctoral researcher who has been running simulations about the evolution of galaxies on the Blue Waters supercomputer at National Center for Supercomputing Applications (NCSA).
“SeedMe provides convenient ways to organize results, which can be easily updated, making sure collaborators or new collaborators all see the most recent results,” added Xu, who has been working with SDSC Director Norman, an astrophysicist by training. “It also avoids the pain of finding figures and plots in emails, and is even proving to be a very useful resource during conference calls.”
Going forward, the SeedMe research team will explore the possibility of making the resource a platform for sharing reusable content. NCSA researcher Matthew Turk is currently investigating how best to share IPython Notebooks from yt software, which is used to analyze and visualize datasets from astrophysical simulations, nuclear engineering, and radio telescope data.
“By building in support to yt for uploading directly to SeedMe, we hope to enable faster, easier, and more seamless sharing of results for anyone using yt,” said Turk. “SeedMe also allows easy control over sharing permissions, while providing an easy-to-use interface for uploading images and data to a central location.”
Tutorials Planned
As an open, web-accessible and searchable research archive, SeedMe is well suited for education and outreach programs. In addition to organizing tutorials, talks, and webinars to train new users and receive user feedback, the program will provide internships to high school and undergraduate students, enabling them to learn about web technologies and the diverse range of research while at the same time support monitoring of content on the project website.
Tutorials and workshops are already in the works, with presentations planned for XSEDE14 and SC14, as well as a webinar as part of SDSC’s Industry Partners program on July 24. Such sessions will show potential users how they can accomplish the following on personal computers and mobile devices directly from compute jobs.
- Get fast feedback on compute jobs
- Share that feedback rapidly with collaborators across the globe
- Provide access to preliminary results to guide further job submission
- Support discussion of feedback and results among collaborators.
Complete details of the SeedMe project can be found at http://www.seedme.org/. The NSF award number for the SeedMe project is OCI-1235505. In addition to Chourasia and Norman, the SeedMe project team includes Mona Wong-Barnum, David Nadeau, and Andrew Ferbert, all with SDSC.
Share This:
You May Also Like
Stay in the Know
Keep up with all the latest from UC San Diego. Subscribe to the newsletter today.


