
New Resource Makes Gene Editing Technology Even More User-Friendly
Published Date
By:
- Liezel Labios
Share This:
Article Content
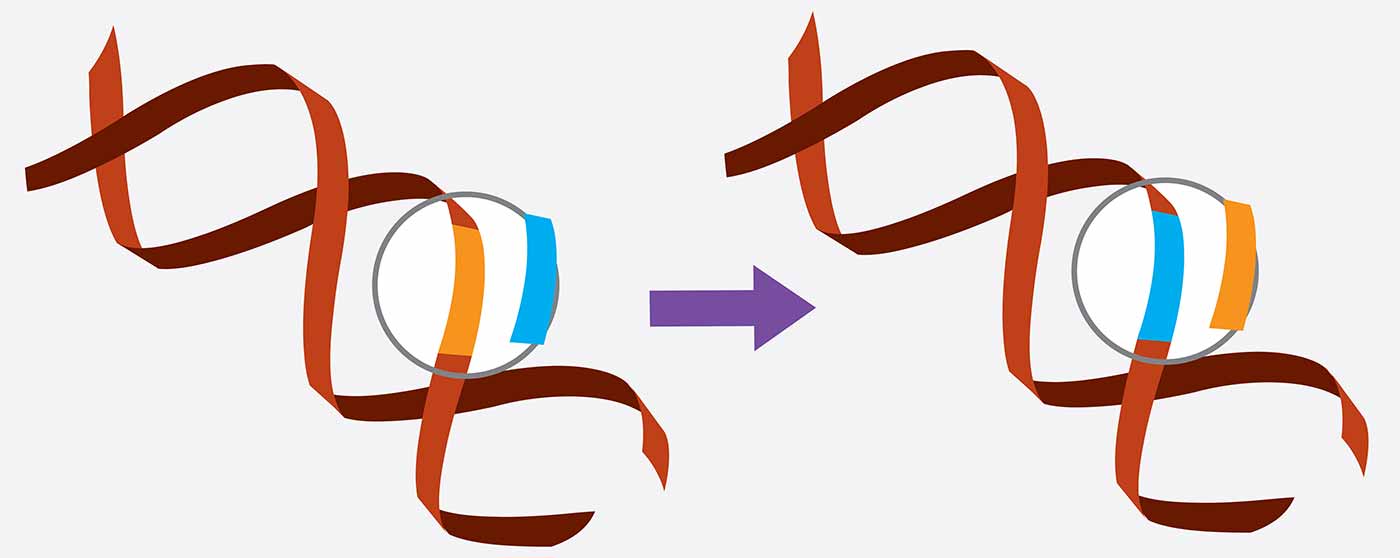
CRISPR/Cas9 is a gene editing tool that can target a particular segment of DNA in living cells and replace it with a new genetic sequence.
Researchers at Harvard University and the University of California, San Diego, have developed a new user-friendly resource to accompany the powerful gene editing tool called CRISPR/Cas9, which has been widely adopted to make precise, targeted changes in DNA. This breakthrough has the potential to facilitate new discoveries in gene therapies and basic genetics research. The research was published in the July 13 issue of Nature Methods.
The study describes an approach to simplify a laborious part of the gene editing process using the CRISPR/Cas9 system: choosing the best components to match specific gene targets.
“We’ve taken a step towards making the CRISPR/Cas9 system more robust,” said Prashant Mali, an assistant professor in the Department of Bioengineering at the UC San Diego Jacobs School of Engineering, and a co-first author of the study.
CRISPR/Cas9 is a relatively new genome engineering tool that can target a particular segment of DNA in living cells — such as a gene mutation — and replace it with a new genetic sequence. This technology ultimately has applications in gene therapies for genetic disorders such as sickle cell anemia and cystic fibrosis.
The CRISPR/Cas9 system has two components: a short “guide RNA” with a sequence matching a particular gene target, and a large protein called Cas9 that cuts DNA precisely at that target. Herein lies the beauty of the CRISPR/Cas9 system: to target another region of the genome, researchers can simply change the guide RNA sequence to match the new gene target.
However, finding the best guide RNA match for a specific gene target is a labor-intensive process. This is because multiple guide RNA sequences can serve as potential matches for each gene target. As a result, researchers might need to test numerous candidates of guide RNAs before finding the most active guide RNA.
Matchmaking software for faster guide RNA selection
To decipher what makes certain guide RNAs better than others, the team conducted what they called a “library-on-library” approach, in which they evaluated a library containing thousands of guide RNAs against a library containing thousands of corresponding gene targets. The team analyzed the data from the library-on-library approach to determine patterns among the guide RNAs that were the most active. Using the data and patterns from these thousands of gene targeting experiments, the team developed a new matchmaking software that predicts and ranks the best guide RNA matches for any given gene target.
“From these experiments, we were able to find features in the guide RNAs that worked and in those that didn’t work. We built a computational model that accounts for all these different features. The end product is an interactive software for users to find guide RNAs that are predicted to be highly specific and highly active for their gene targets,” said Raj Chari, a research fellow working in the lab of Professor George Church in the Department of Genetics at Harvard Medical School, and a co-first author of the study.
“We hope to minimize the time and work in finding the most successful guide RNA sequence for a gene target, which will be helpful for finding new gene therapies,” said Chari.
“Overall, this new method offers a simple approach to assess a large number of guide RNAs in a short amount of time. We believe this will be a useful resource for the community towards designing improved genome engineering experiments,” added Mali.
This work was supported by the National Institutes of Health (grant P50HG005550), a Banting Fellowship from the Canadian Institutes of Research, University of California San Diego startup funds, and a Burroughs Wellcome Career Award.
The team was led by George Church, HMS Robert Winthrop Professor of Genetics at Harvard Medical School. Mark Moosburner, a Ph.D. student in the Scripps Institute of Oceanography at UC San Diego, was also a member of the research team.
Share This:
You May Also Like
Stay in the Know
Keep up with all the latest from UC San Diego. Subscribe to the newsletter today.


