Scientists Discover a New Signaling Pathway and Design a Novel Drug for Liver Fibrosis
Health & Behavior
Navigated and unguided modes of populatiom expansion. The radial density profile is shown for the navigated expansion process taken by chemotactic bacteria (top panel) and unguided expansion process taken by motile but non-chemotactic bacteria (bottom panel). Each is inoculated at the center of a soft-agar plate at time t=0. The density bulge at the leading front of navigated expansion is the feature seen distinctively in the photos of Fig. 1A as the “swarm ring.”
One of the best studied topics of molecular biology is bacterial chemotaxis—the movement of bacterial cells in response to chemical stimuli. While scientists at UC San Diego thought they—and the science community in general—knew everything there was to know about how and why bacterial cells moved around, they were surprised to realize how little they understood about how bacteria moved around as a group. New findings on this latest research are published as rare back-to-back articles in Nature, Nov. 6, 2019.
In the first paper, Distinguished Professor of Physics and Biology Terence Hwa, along with Postdoctoral Fellow Jonas Cremer and other UC San Diego researchers, worked with the model bacterium E. coli to show that chemotaxis provides a mechanism for bacterial populations to undertake a highly effective mode of organized exploration into unoccupied surrounding territories. Instead of initiating exploration when nutrients ran out in the current environment as commonly believed, the research team found that bacteria launched exploration when growing happily in their current environment.
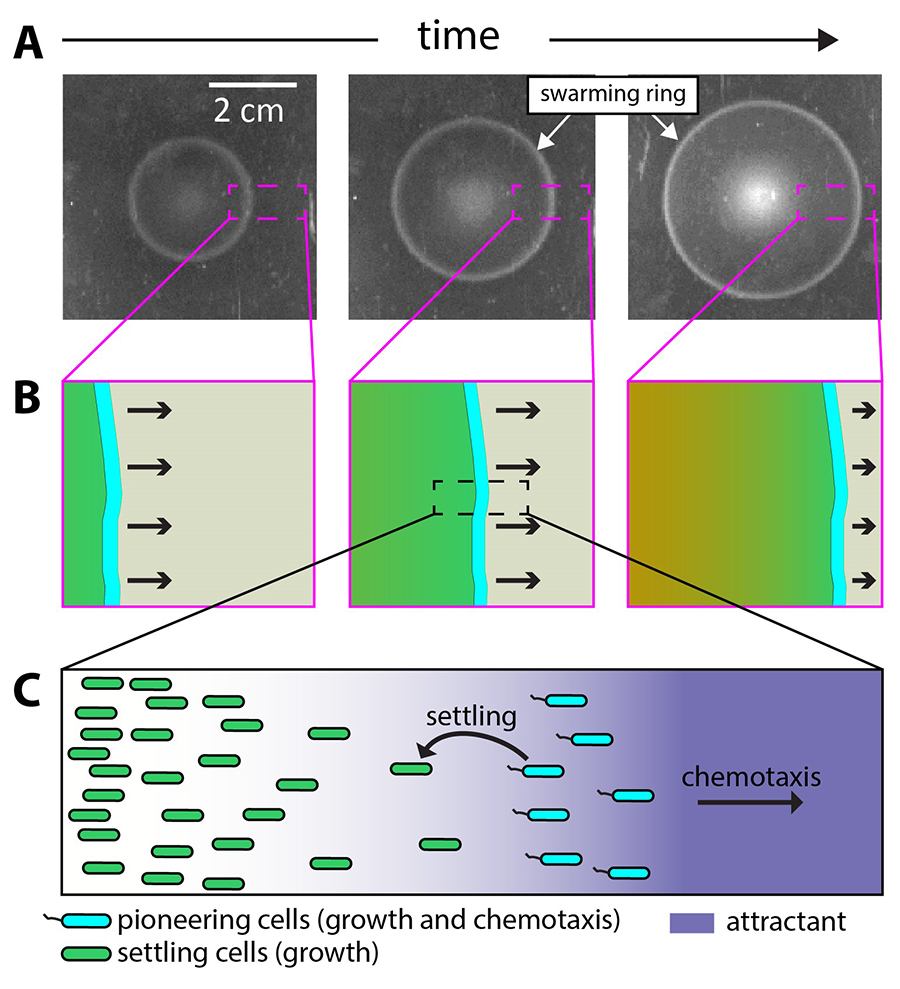
Figure 1: Expanding population of chemotactic bacteria. A) Inoculating a few E. coli cells at the middle of a nutrientre-rich soft-agar plate results in the formation of a “swarm ring,” which would expand across the plate, reaching the edge in ~12 hours. B) and C) The swarm ring is comprised of a dense region of pioneering cells (cyan) which uses chemotaxis to move up the gradient of chemoattracant (purple). A small fraction of pioneering cells would fall behind the swarm ring to settle and replicate in their locality (indicated as green cells), eventually reaching saturating densities (brown region in panel B).
“In fact, the better the environment, the more actively engaged are the bacteria in exploration—reminiscent of how well-doing corporations such as Google® engage more actively in diversification, despite their booming core businesses,” said Hwa.
Using high-speed microscopes, the researchers were able to track the motion of individual bacteria in a population of many thousands that formed the so-called “swarm ring” of chemotaxis—an easy-to-see “ripple” that expands outward after a drop of bacteria is introduced to a nutrient-rich habitat (see Figure 1A). To their surprise, the researchers observed that the “swarm ring” was not merely a small group of distracted bacteria as commonly thought. Instead, the ring of bacteria included pioneering explorers that replicated as they migrated, leaving behind a trail of off-spring (see Figure 1B,C).
“Unlike ant trails, the trailing bacteria would not migrate, but would replicate in their respective localities behind the expanding ring, eventually filling the entire space enclosed by the ring,” explained Hwa. “These bacteria appear to be smart enough to know that exponential growth cannot be sustained in a given locality for long. So pioneering explorers are sent out while nutrients are plentiful, in order for the population not to be stuck when nutrients run out.”
Hwa noted that while scientists knew well that the bacteria navigated their way around by moving toward higher concentrations of certain molecules called “chemoattractants,” it was long thought that these molecules were “attractive” because they were good nutrient sources. Hwa’s team, however, showed that potent chemoattractants were not good nutrient sources for the bacteria, and they were most potent when presented in low doses accompanied by good nutrients. This led the researchers to realize that the chemoattractants are more like aroma than food to the bacteria (see Figure 1C).
“We showed how low levels of chemoattractants can be effectively shaped by the pioneering bacteria into a gradient to navigate the exploration process, establishing the direction and also enhancing the speed of population movement,” said Cremer.
The research team concluded their work by comparing the bacteria’s navigated mode of exploration, settlement and replication to the unguided “range expansion” process not involving chemotaxis as commonly assumed in ecology (see Movie 1). As the population spreads faster and yields larger gains under navigated exploration, Hwa and Cremer predict this process may be employed widely in nature, well beyond the bacterial world.
“It is not hard to imagine that a population of higher animals may use a similar strategy as the bacteria—modifying environment and generating cues that direct the population to expand rapidly into unoccupied territories.”
The second paper published in Nature builds on lessons learned from the first one. In this paper, Hwa and Cremer, working with scientists based at the Chinese Academy of Sciences in Shenzhen, China, studied the evolutionary underpinnings of the navigated exploration strategy undertaken by chemotactic bacteria.
Through a series of laboratory evolution experiments selecting for E. coli that successfully colonize habitats of different sizes, the researchers studied the fitness advantages of chemotactic bacteria at different distances from a common starting position (see Figure 2A,B).
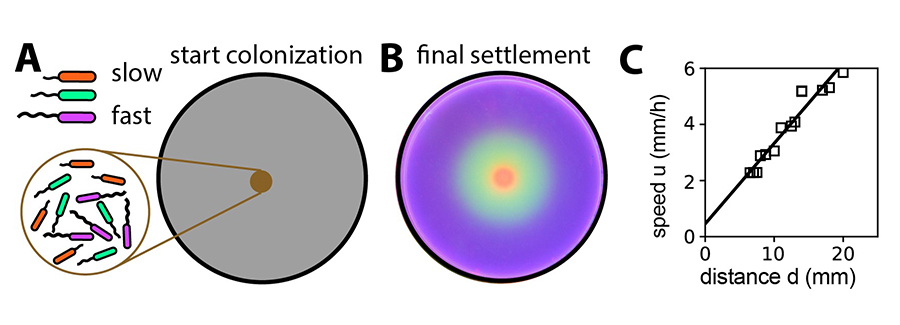
Figure 2: Successful colonization of different habitats depend on the population’s expansion speed and growth rate. A) An equal mixture of slow and fast swimming bacteria (indicated by different colors) was inoculated at the center of the soft-agar plate. B) After the mixed population had ample time to expand and grow to high densities, the slowest strain (red) dominated at small distances, the fastest strain dominated at the distances far away, with the strain with intermediate speed dominating the intermediate distances. C) Detailed evolution experiments selecting for strains most successful in colonizing different radial distances (d) found the optimal expansion speed (u) to be linear with the selection distance as predicted by the authors’ mathematical analysis.
“For multiple types of bacteria with a range of speeds, traditional wisdom would expect the first species reaching an unoccupied habitat to have a decisive advantage in colonizing that habitat, compared to other bacteria with similar growth behavior,” said Hwa. “But our evolution experiments showed surprisingly that faster is not necessary advantageous. While faster strains have advantages at distances far away, slower strains are advantageous at smaller distances.”
These findings, together with results from the first paper, allowed the team to discover a constraint between the expansion speed of a population and the size of the habitat that the population can successfully colonize—it is simply the size that the expanding population front can reach in several replication time.
Hwa explained that this constraint reflects a required coordination between the speed of expansion and the rate of population growth needed to successfully colonize newly gained habitats.
“If expansion should occur at a speed faster than the organisms’ means to replicate in the new land, then they would end up losing their claim to the land even if they occupied it first,” said Hwa. “This is perhaps analogous to why Genghis Khan’s army could sweep across half of the civilized world while unable to rule the conquered land.”
The proposed constraint was quantitatively validated by the researchers, who repeated the evolution experiments in different habitats, allowing for many combinations of speeds and growth rates (see Figure 2C). According to Hwa, quantitatively predicting the outcome of evolution experiments is very rare, but the success of these studies may offer hope for more effective approaches to investigating evolutionary processes in the future.
Hwa also suggested that the kind of constraint they uncovered on exploration strategy might apply to other types of exploration systems, including perhaps growing tumors or even expanding businesses.
Coauthors of the first paper include Tomoya Honda of the Division of Biological Sciences and Ying Tang, Jerome Wong-Ng and Massimo Vergassola of the Department of Physics. On the second paper, Hwa and Cremer worked with coauthors Weirong Liu, Dengjin Li and Chenli Liu from the Chinese Academy of Sciences. The research at UC San Diego was supported by the National Institutes of Health (grant no. R01GM95903), the NSF Program PoLS (grant no. 1411313) and a JASSO long-term graduate fellowship award.
Keep up with all the latest from UC San Diego. Subscribe to the newsletter today.